PCAngsd
PCAngsd is a program that estimates the covariance matrix and individual allele frequencies for low-depth next-generation sequencing (NGS) data in structured/heterogeneous populations using principal component analysis (PCA) to perform multiple population genetic analyses using genotype likelihoods. Since version 0.98, PCAngsd was re-written to be based on Cython for computational bottlenecks and parallelization.
The main method was published in 2018 and can be found here: [1]
The HWE test was published in 2019 and can be found here: [2]
Overview
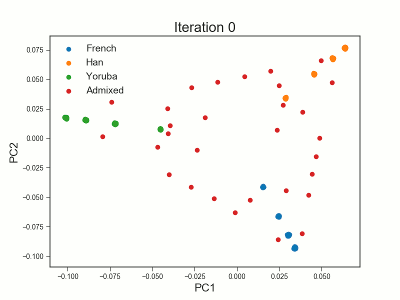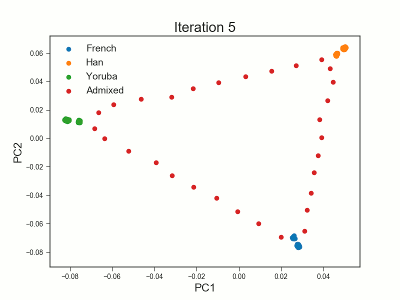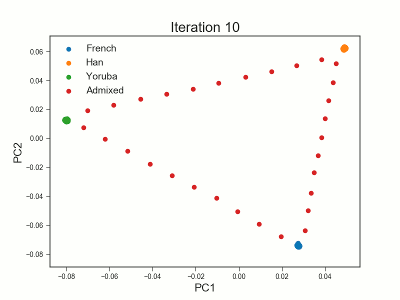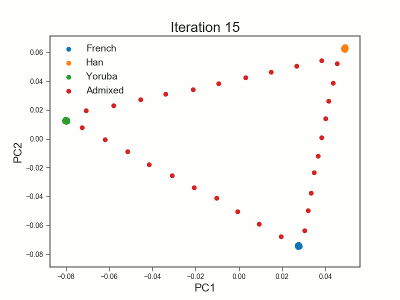
Framework for analyzing low-depth next-generation sequencing (NGS) data in heterogeneous/structured populations using principal component analysis (PCA). Population structure is inferred by estimating individual allele frequencies in an iterative approach using a truncated SVD model. The covariance matrix is estimated using the estimated individual allele frequencies as prior information for the unobserved genotypes in low-depth NGS data.
The estimated individual allele frequencies can further be used to account for population structure in other probabilistic methods. PCAngsd can perform the following analyses:
- Covariance matrix
- Admixture estimations
- Inbreeding coefficients (both per-individual and per-site)
- HWE test
- Genome-wide selection scan
- Genotype calling
- Estimate NJ tree of samples
Older versions of PCAngsd can be found here [3].
Download and Installation
PCAngsd should work on all platforms meeting the requirements but server-side usage is highly recommended. Installation has only been tested on Linux systems.
Get PCAngsd and build
git clone https://github.com/Rosemeis/pcangsd.git cd pcangsd/ python setup.py build_ext --inplace
Install dependencies:
The required set of Python packages are easily installed using the pip command and the 'requirements.txt file' included in the 'pcangsd' folder.
pip install --user -r requirements.txt
Quick start
PCAngsd is used by running the main caller file pcangsd.py. To see all available options use the following command:
python pcangsd.py -h # Genotype likelihoods using 64 threads python pcangsd.py -beagle input.beagle.gz -out output -threads 64 # PLINK files (using file-prefix, *.bed, *.bim, *.fam) python pcangsd.py -beagle input.plink -out output -threads 64
PCAngsd accepts either genotype likelihoods in Beagle format or PLINK genotype files. Beagle files can be generated from BAM files using ANGSD. For inference of population structure in genotype data with non-random missigness, we recommend our EMU software that performs accelerated EM-PCA, however with fewer functionalities than PCAngsd (#soon).
PCAngsd will mostly output files in binary Numpy format (.npy) with a few exceptions. In order to read files in python:
import numpy as np
C = np.genfromtxt("output.cov") # Reads in estimated covariance matrix (text)
D = np.load("output.selection.npy") # Reads PC based selection statistics
R can also read Numpy matrices using the "RcppCNPy" R library:
library(RcppCNPy)
C <- as.matrix(read.table("output.cov")) # Reads in estimated covariance matrix
D <- npyLoad("output.selection.npy") # Reads PC based selection statistics
An example of generating genotype likelihoods in ANGSD and output them in the required Beagle text format.
./angsd -GL 2 -out input -nThreads 4 -doGlf 2 -doMajorMinor 1 -doMaf 2 -SNP_pval 1e-6 -bam bam.filelist
Tutorial
Please refer to the tutorial's page [4]
Options
# See all options in PCAngsd python pcangsd.py -h
General usage
- -beagle [Beagle file]
Input file of genotype likelihoods in Beagle format (.beagle.gz).
- -filter [Text file]
Input file of 1's or 0's whether to keep individuals or not.
- -plink [Prefix for binary PLINK files]
Path to PLINK files using their ONLY prefix (.bed, .bim, .fam).
- -plink_error [float]
Incorporate errors into genotypes by specifying rate as argument.
- -minMaf [float]
Minimum minor allele frequency threshold. (Default: 0.05)
- -maf_iter [int]
Maximum number of EM iterations for computing the population allele frequencies (Default: 200).
- -maf_tole [float]
Tolerance value in EM algorithm for population allele frequencies estimation (Default: 1e-4).
- -iter [int]
Maximum number of iterations for estimation of individual allele frequencies (Default: 100).
- -tole [float]
Tolerance value for update in estimation of individual allele frequencies (Default: 1e-5).
- -hwe [.lrt.npy file]
Input file of LRT binary file from previous PCAngsd run to filter based on HWE.
- -hwe_tole [float]
Threshold for HWE filtering of sites.
- -e [int]
Manually select the number of eigenvalues to use in the modelling of individual allele frequencies (Default: Automatically tested using MAP test).
- -pi [.pi.npy file]
Load previous estimation of individual allele frequencies to skip covariance estimation.
- -maf_save
Choose to save estimated population allele frequencies (Binary). Numpy format (.npy).
- -pi_save
Choose to save estimated individual allele frequencies (Binary). Numpy format (.npy). Can be used with the '-pi' command.
- -dosage_save
Choose to save estimated genotype dosages (Binary). Numpy format (.npy).
- -post_save
Choose to save the posterior genotype probabilities. Beagle format (.beagle).
- -sites_save
Choose to save the kept sites after filtering which is useful for downstream analysis. Outputs a file of 1's and 0's for keeping a site or not, respectively.
- -threads [int]
Specify the number of thread(s) to use (Default: 1).
- -out [output prefix]
Fileprefix for all output files created by PCAngsd (Default: "pcangsd").
Selection
Perform PC-based genome-wide selection scans using posterior expectations of the genotypes (genotype dosages):
- -selection
Using an extended model of FastPCA. Performs a genome-wide selection scan along all significant PCs. Outputs the selection statistics and must be converted to p-values by user. Each column reflect the selection statistics along a tested PC and they are χ²-distributed with 1 degree of freedom.
- -pcadapt
Using an extended model of pcadapt. Performs a genome-wide selection scan across all significant PCs. Outputs the z-scores and must be converted to test statistics with the provided script 'pcangsd/scripts/pcadapt.R', and the test statistics are χ²-distributed with K degree of freedom.
- -snp_weights
Output the SNP weights of the significant K eigenvectors.
Inbreeding
- -inbreedSites
Estimate per-site inbreeding coefficients accounting for population structure and perform likehood ratio test for detecting sites deviating from HWE [5].
- -inbreedSamples
Estimate per-individual inbreeding coefficients accounting for population structure which is based on an extension of ngsF for structured populations.
- -inbreed_iter [int]
Maximum number of iterations for inbreeding EM algorithm. (Default: 200)
- -inbreed_tole [float]
Tolerance value for inbreeding EM algorithm in estimating inbreeding coefficients. (Default: 1e-4)
Call genotypes
Genotypes can be called from posterior genotype probabilities by incorporating the individual allele frequencies as prior information.
- -geno [float]
Call genotypes with defined threshold.
- -genoInbreed [float]
Call genotypes with defined threshold also taking inbreeding into account. '-inbreedSamples' must also be called for using this option.
Admixture
Individual admixture proportions and ancestral allele frequencies can be estimated assuming K ancestral populations using an accelerated mini-batch NMF method.
- -admix
Toggles admixture estimations. Estimates admixture proportions and ancestral allele frequencies.
- -admix_K [int]
Not recommended. Override the number of ancestry components (K) to use, instead of using K=e-1.
- -admix_iter [int]
Maximum number of iterations for admixture estimations using NMF. (Default: 200)
- -admix_tole [float]
Tolerance value for update in admixture estimations using NMF. (Default: 1e-5)
- -admix_alpha [float
Specify alpha (sparseness regularization parameter). (Default: 0)
- -admix_auto [float]
Enable automatic search for optimal alpha using likelihood measure, by giving soft upper search bound of alpha.
- -admix_seed [int]
Specify seed for random initializations of factor matrices in admixture estimations.
Tree
- -tree
Construct neighbour-joining tree of samples from estimated covariance matrix estimated based on indivdual allele frequencies.
- -tree_samples
Provide a list of sample names of all individuals to construct a beautiful tree.
Citation
Our methods for inferring population structure have been published in GENETICS:
Inferring Population Structure and Admixture Proportions in Low Depth NGS Data
Our method for testing for HWE in structured populations has been published in Molecular Ecology Resources: