RelateAdmix
Brief description
This page contains information about the program called relateAdmix, which can be used to infer relatedness coefficients for pairs of individuals or inbreeding coefficients even if they are admixed. The program has both an R interface and a C interface. Below is a description of how to install and use each of them. To be able to infer the relatedness you will need to know the individuals admixture proportions and the allele frequencies in each of the possible populations. This can be done e.g. using the program ADMIXTURE as shown in the example of how to use the C interface.
Installation - Linux/MAC
Both the C and R versions of the program have been made for linux but should work on other unix like systems. You need to have zlib installed. Some MAC users might also have to use makevars
Windows is not supported
Download
I move all the code to github [1]
Installation of R package
If you have the devtools packages (https://github.com/hadley/devtools) installed in R then you can install the package i R directly from github
library(devtools)
install_github("aalbrechtsen/relateAdmix")
Or install from the command line
git clone https://github.com/aalbrechtsen/relateAdmix.git R CMD INSTALL relateAdmix
or build and install
git clone https://github.com/aalbrechtsen/relateAdmix.git R CMD build relate R CMD INSTALL Relate_<add version number>.tar.gz
Problems with zlib
If your R installation for some reason does not link to your zlib (e.i. gzopen missing) or you have a really old zlib installation then you can install the R part using
rm relateAdmix/src/Cinterface.* rm relateAdmix/src/Makefile #only needed if you compiled the C program R CMD INSTALL relateAdmix
Installation of C/C++ program
git clone https://github.com/aalbrechtsen/relateAdmix.git cd relateAdmix/src cp CPP_Makefile Makefile make
Run example
Run example using R
After installing the package you can load it into R and try the example
library(relateAdmix) example(relate)
This shows an example of how to use the package. More information can be found in the man pages
?relate
Run example from terminal
After installing the C version of the program you can try running it on the example data set in the data folder, which consists of 104290 SNP for 126 individuals that are admixed from 2 source populations.
If you are in the src folder where you installed relateAdmix and you have the software ADMIXTURE installed this can be done as follows:
cd ../data # First run Admixture using a plink ".bed" as input to produce population specific allele # frequencies (smallPlink.2.P) and individual ancestry proportions (smallPlink.2.Q). # (note other programs can be used instead of Admixture, e.g. Structure and FRAPPE) admixture smallPlink.bed 2 # Then run relateAdmix with plink bed, bim and fam files plus the Admixture output files as input ../src/relateAdmix -plink smallPlink -f smallPlink.2.P -q smallPlink.2.Q -P 20
NB!. Only use binary plink (.bed) since ADMIXTURE switches allele frequencies when using .ped files
Quick R pdf plot from the commandline:
# Plot the results in R (R needs to be installed)
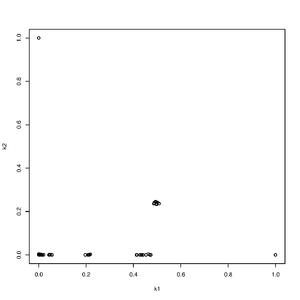
Rscript -e "r<-read.table('output.k',head=T,as.is=T);pdf('rel.pdf');plot(r[,4],r[,5],ylab='k2',xlab='k1');dev.off()"
Options
see options by running with out additional arguments:
../src/relateAdmix Arguments: -plink name of the binary plink file (excluding the .bed) -fname Ancestral population frequencies -qname Admixture proportions -o name of the output file Setup: -P Number of threads -F 1 if you want to estimate inbreeding
- -plink [FILE]
The binary plink file without postfix
- -f/-fname [FILE]
Ancestral population frequencies (e.g. estimated from ADMIXTURE with the .P filename
- -q/-qname [FILE]
Ancestral admxiture proportions (e.g. estimated from ADMIXTURE with the .Q filename
- -o [NAME]
Output filename
- -P [INTEGER]
Number of threads (not implemented for inbreeding)
- -F [0 or 1]
Use -F 1 if you want to estimate the inbreeding coefficient instead of relatedness
Output file format
Example of output
ind1 ind2 k0 k1 k2 nIter 0 1 0.999941 0.000038 0.000021 26 0 2 0.999979 0.000010 0.000011 29 0 3 0.999953 0.000029 0.000018 26 0 4 0.999952 0.000023 0.000025 26 0 5 0.999972 0.000020 0.000007 26 0 6 0.999995 0.000003 0.000002 26 0 7 0.999995 0.000003 0.000002 26 0 8 0.999894 0.000069 0.000038 32 0 9 0.999894 0.000069 0.000038 32 0 10 0.999903 0.000071 0.000026 26 0 11 0.999903 0.000071 0.000026 26
The first two columns are the individuals number. The next three columns are the estimated relatedness coefficients and the last column is the number of iterations used.
Input file format
The input consists of three files describignt the genotype data, a file with admixture proportions for each individual and a file with allele frequencies for each SNP for each source population. The genotype data files are plink bed/bim/fam files. And the remaining two files are in the output format for the program ADMIXTURE:
Example of the content of an admixture proportion file (for 3 populations)
0.531631 0.468359 0.000010 0.564461 0.435529 0.000010 0.850660 0.149330 0.000010 0.630527 0.369463 0.000010 0.747429 0.219346 0.033225 0.999980 0.000010 0.000010 0.999980 0.000010 0.000010 0.682072 0.317918 0.000010 0.000010 0.999980 0.000010 0.793133 0.206857 0.000010
Each row is an individual and each column is a population. The admixture proportions for each individual must sum to 1
Example of the allele frequency file (for 3 populations)
0.312722 0.208605 0.999990 0.881352 0.999990 0.966966 0.708206 0.838869 0.932119 0.427789 0.620694 0.532966 0.411998 0.622253 0.534072 0.427789 0.620694 0.532966 0.440817 0.581630 0.618751 0.733733 0.985281 0.953523 0.724083 0.451452 0.784607 0.811161 0.578612 0.787782
Each row is an SNP and each column is a population. When using plink files the allele frequency is the MAJOR allele frequency.
Citing and references
relateAdmix
Moltke, I, Albrechtsen, A (2013). RelateAdmix: a software tool for estimating relatedness between admixed individuals. Bioinformatics. pubmed bibtex
Inbreeding
The model used for inbreeding is described in Ida Moltke, Matteo Fumagalli, Thorfinn S Korneliussen, Jacob E Crawford, Peter Bjerregaard, Marit E Jørgensen, Niels Grarup, Hans Christian Gulløv, Allan Linneberg, Oluf Pedersen, Torben Hansen, Rasmus Nielsen, Anders Albrechtsen Uncovering the genetic history of the present-day Greenlandic population. Am. J. Hum. Genet.: 2015, 96(1);54-69 Article
ADMIXTURE
D.H. Alexander, J. Novembre, and K. Lange. Fast model-based estimation of ancestry in unrelated individuals. Genome Research, 19:1655–1664, 2009.
change log
- 0.14 made more MAC usable (I think). Thanks to Paul Lott for reporting it and for suggestions and Thorfinn Sand for changing it
- 0.13 added extra check for file exists to give instant errors + changes all printf to fprintf(stderr,
- 0.11 changed threading to a fixed pool of threads
- 0.10 optimized code
- 0.09 added error for when the number of sites and individuals does not match between files
- 0.08 fixed a bug that would sometimes print an extra line when multiple threaded
- 0.07 fixed a small leak