NgsAdmixv2: Difference between revisions
| (39 intermediate revisions by 2 users not shown) | |||
| Line 24: | Line 24: | ||
NGSadmix can be installed independently or as a part of ANGSD. | NGSadmix can be installed independently or as a part of ANGSD. | ||
===NGSadmix Independent Installation=== | ====NGSadmix Independent Installation==== | ||
1. Login to your server using ssh on your terminal window. | 1. Login to your server using ssh on your terminal window. | ||
| Line 38: | Line 38: | ||
:<code>g++ ngsadmix32.cpp -O3 -lpthread -lz -o NGSadmix</code> | :<code>g++ ngsadmix32.cpp -O3 -lpthread -lz -o NGSadmix</code> | ||
====NGSadmix Installation from ANGSD==== | |||
:NGSadmix is part of the package ANGSD. To install ANGSD, please follow the instructions here [http://popgen.dk/angsd/index.php/Installation] | |||
<pre> | ==Parameters== | ||
</pre> | |||
All parameters are set using '''-par value'''. | |||
For example, to get additional information, you would write '''-printInfo 1'''. | |||
<pre>./NGSadmix </pre> | |||
Arguments: | |||
::'''-likes''' .beagle format filename with genotype likelihoods | |||
::'''-K''' Number of ancestral populations | |||
Optional: | |||
: | |||
::'''-fname''' Ancestral population frequencies | |||
:: | |||
:: | ::'''-qname''' Admixture proportions | ||
::'''-outfiles''' Prefix for output files | |||
::'''-printInfo''' print ID and mean maximum allele frequency (maf) for the SNPs that were analysed | |||
Setup: | |||
::'''-seed''' Seed for initial guess in EM algorithm (a number lower than 1M is preferred). | |||
:: The same seed can be used to reproduce the analysis, and 3 different seeds can be used to test convergence. | |||
::'''-P''' Number of threads | |||
::'''-method''' 0 indicates no acceleration of EM algorithm. Please refer to the paper for more information. | |||
::'''-misTol''' Tolerance for considering a site as missing. Default = 0.05. | |||
:: To include high quality genotypes only, increase this value (for example, 0.9) | |||
Stop criteria: | Stop criteria: | ||
</ | ::'''-tolLike50''' Loglikelihood difference in 50 iterations. Default= 0.1 | ||
::'''-tol''' Tolerance for convergence. Default = 1x10<sup>-5</sup>. Use maller values for higher accuracy. | |||
:: It's the maximum squared difference of F and Q (please refer to the paper for formula). | |||
::'''-dymBound''' Use dymamic boundaries (1: yes (default) 0: no). | |||
::'''-maxiter''' Maximum number of EM iterations. Default = 2000 (high value). | |||
:: In case it doesn't converge, this value needs to be higher. | |||
Filtering: | |||
::'''-minMaf''' Minimum minor allele frequency. Default = 5% | |||
::'''-minLrt''' Minimum likelihood ratio value for maf>0. Default = 0 | |||
::'''-minInd''' Minumum number of informative individuals. Default = 0 | |||
:: It only keeps sites where there is at least x # of individuals with NGS data. | |||
==Input File== | ==Input File== | ||
| Line 105: | Line 103: | ||
The input file contains genotype likelihoods in a .beagle file format [http://faculty.washington.edu/browning/beagle/beagle.html]. | The input file contains genotype likelihoods in a .beagle file format [http://faculty.washington.edu/browning/beagle/beagle.html]. | ||
and can be compressed with gzip. | and can be compressed with gzip. | ||
=== BAM files === | |||
If you have BAM files you can use [[ANGSD]] to produce genotype likelihoods in .beagle format. Please | |||
see [http://www.popgen.dk/angsd/index.php/Beagle_input Creation of Beagle files with ANGSD] | |||
=== VCF files === | |||
If you already have made a VCF file that contains genotype likehood information then it should be possible to convert .vcf files with genotype likelihoods to .beagle file via vcftools [https://vcftools.github.io/man_latest.html] | |||
<pre> | |||
vcftools --vcf input.vcf --out test --BEAGLE-GL --chr 1,2 | |||
</pre> | |||
Chromosome has to be specified. | |||
You can also use bcftools' [https://samtools.github.io/bcftools/bcftools.html] 'query' option for generating a .beagle file from a .vcf file. | |||
==Output Files== | ==Output Files== | ||
| Line 114: | Line 122: | ||
* Log likelihood of the estimates: a .log file that summarizes the run. The Command line used for running the program, what the likelihood is every 50 iterations, and finally how long it took to do the run. | * Log likelihood of the estimates: a .log file that summarizes the run. The Command line used for running the program, what the likelihood is every 50 iterations, and finally how long it took to do the run. | ||
* Estimated allele frequency: a zipped .fopt file, that contains an estimate of the allele frequency in each of the 3 assumed ancestral populations | * Estimated allele frequency: a zipped .fopt file, that contains an estimate of the allele frequency in each of the 3 assumed ancestral populations. There is a line for each locus. | ||
* Estimated admixture proportions: a .qopt file, that contains an estimate of the individual's ancestry proportion (admixture) from each of the three assumed ancestral populations for all individuals | * Estimated admixture proportions: a .qopt file, that contains an estimate of the individual's ancestry proportion (admixture) from each of the three assumed ancestral populations for all individuals. There is a line for each individual. | ||
==Run command example== | |||
Download the input file, for example | |||
::<code>wget popgen.dk/software/download/NGSadmix/data/input.gz</code> | |||
Execute NGSadmix | |||
::<code>./NGSadmix -likes input.gz -K 3 -P 4 -o myoutfiles -minMaf 0.05</code> | |||
::* '''-likes''' Input .beagle file of genotype likelihoods = input.gz | |||
::* '''-K''' Ancestral populations K=3 | |||
::* '''-P''' Number of threads used, as computer cores = 4 | |||
::* '''-o''' Output prefix, output file names = myoutfiles | |||
::*'''-minMaf''' SNPs with MAF > 5% = 0.05 | |||
===Detailed Examples and Tutorial=== | |||
Please refer to the tutorial's page [http://www.popgen.dk/software/index.php/NgsAdmixTutorial] | |||
==Citation== | ==Citation== | ||
Latest revision as of 10:44, 12 July 2019
This page contains information about the program called NGSadmix, which is a very nice tool for estimating individual admixture proportions from NGS data. It is based on genotype likelihoods and works well for medium and low coverage NGS data. It is a fancy multithreaded c/c++ program which makes it useful for large datasets.
The great thing about NGSadmix is that it is a new method that takes the uncertainty introduced in NGS sequencing data into account when inferring an individual's ancestry by using genotype likelihoods that considers the uncertainty caused by unobserved genotypes.
As with the other existing software, ADMIXTURE and STRUCTURE, NGSadmix is only sensitive to admixture recent enough to cause structures in the population in terms of differing allele frequencies. Historical admixture events after which many generations has passed in the population, leaves no signature in terms of systematic differences in allele frequencies between individuals and are not a concern in association studies.
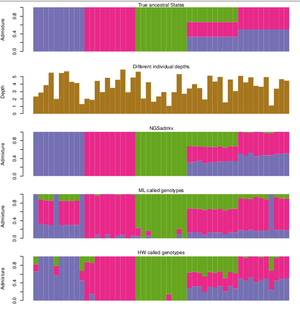
The method was published in 2013 and can be found here: [1]
Software Download
The latest version of NGSadmix is ngsadmix32 from June 25, 2013 and can be downloaded here: [2].
- Older Versions
- The previous version of NGSadmix, ngsadmix31 can be found here: [3].
- Version Log:
- v32 june 25-2013; modified code such that it now compiles on OSX
- v31 june 24-2013; First public version.
Installation
NGSadmix can be installed independently or as a part of ANGSD.
NGSadmix Independent Installation
1. Login to your server using ssh on your terminal window.
2. Create the directory where you will install your software and enter it, such as
mkdir ~/Softwarecd ~/Software
3. Download the source code:
4. Configure, Compile and Install:
g++ ngsadmix32.cpp -O3 -lpthread -lz -o NGSadmix
NGSadmix Installation from ANGSD
- NGSadmix is part of the package ANGSD. To install ANGSD, please follow the instructions here [4]
Parameters
All parameters are set using -par value. For example, to get additional information, you would write -printInfo 1.
./NGSadmix
Arguments:
- -likes .beagle format filename with genotype likelihoods
- -K Number of ancestral populations
Optional:
- -fname Ancestral population frequencies
- -qname Admixture proportions
- -outfiles Prefix for output files
- -printInfo print ID and mean maximum allele frequency (maf) for the SNPs that were analysed
Setup:
- -seed Seed for initial guess in EM algorithm (a number lower than 1M is preferred).
- The same seed can be used to reproduce the analysis, and 3 different seeds can be used to test convergence.
- -P Number of threads
- -method 0 indicates no acceleration of EM algorithm. Please refer to the paper for more information.
- -misTol Tolerance for considering a site as missing. Default = 0.05.
- To include high quality genotypes only, increase this value (for example, 0.9)
Stop criteria:
- -tolLike50 Loglikelihood difference in 50 iterations. Default= 0.1
- -tol Tolerance for convergence. Default = 1x10-5. Use maller values for higher accuracy.
- It's the maximum squared difference of F and Q (please refer to the paper for formula).
- -dymBound Use dymamic boundaries (1: yes (default) 0: no).
- -maxiter Maximum number of EM iterations. Default = 2000 (high value).
- In case it doesn't converge, this value needs to be higher.
Filtering:
- -minMaf Minimum minor allele frequency. Default = 5%
- -minLrt Minimum likelihood ratio value for maf>0. Default = 0
- -minInd Minumum number of informative individuals. Default = 0
- It only keeps sites where there is at least x # of individuals with NGS data.
Input File
The input file contains genotype likelihoods in a .beagle file format [5]. and can be compressed with gzip.
BAM files
If you have BAM files you can use ANGSD to produce genotype likelihoods in .beagle format. Please see Creation of Beagle files with ANGSD
VCF files
If you already have made a VCF file that contains genotype likehood information then it should be possible to convert .vcf files with genotype likelihoods to .beagle file via vcftools [6]
vcftools --vcf input.vcf --out test --BEAGLE-GL --chr 1,2
Chromosome has to be specified.
You can also use bcftools' [7] 'query' option for generating a .beagle file from a .vcf file.
Output Files
The analysis performed by NGSadmix produces 4 files:
- Log likelihood of the estimates: a .log file that summarizes the run. The Command line used for running the program, what the likelihood is every 50 iterations, and finally how long it took to do the run.
- Estimated allele frequency: a zipped .fopt file, that contains an estimate of the allele frequency in each of the 3 assumed ancestral populations. There is a line for each locus.
- Estimated admixture proportions: a .qopt file, that contains an estimate of the individual's ancestry proportion (admixture) from each of the three assumed ancestral populations for all individuals. There is a line for each individual.
Run command example
Download the input file, for example
wget popgen.dk/software/download/NGSadmix/data/input.gz
Execute NGSadmix
./NGSadmix -likes input.gz -K 3 -P 4 -o myoutfiles -minMaf 0.05
- -likes Input .beagle file of genotype likelihoods = input.gz
- -K Ancestral populations K=3
- -P Number of threads used, as computer cores = 4
- -o Output prefix, output file names = myoutfiles
- -minMaf SNPs with MAF > 5% = 0.05
Detailed Examples and Tutorial
Please refer to the tutorial's page [8]
Citation
http://www.genetics.org/content/early/2013/09/03/genetics.113.154138.full.pdf
Skotte, L., Korneliussen, T. S., & Albrechtsen, A. (2013). Estimating individual admixture proportions from next generation sequencing data. Genetics, 195(3), 693–702. doi:10.1534/genetics.113.154138
- Bibtex
- % 24026093
- @Article{pmid24026093,
- Author="Skotte, L. and Korneliussen, T. S. and Albrechtsen, A. ",
- Title="{{E}stimating {I}ndividual {A}dmixture {P}roportions from {N}ext {G}eneration {S}equencing {D}ata}",
- Journal="Genetics",
- Year="2013",
- Pages=" ",
- Month="Sep"
- }